
Studies on a landscape of perfluoroaromatic-reactive peptides
Studies on a landscape of perfluoroaromatic-reactive peptides
Org. Biomol. Chem., 2019,17, 1862-1868
Ethan D. Evans and Bradley L. Pentelute
Abstract
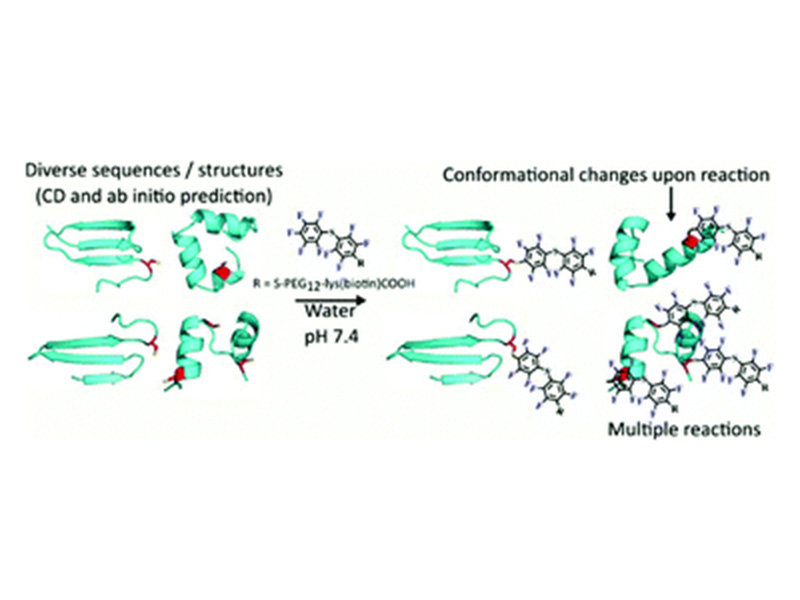
We investigated 26 midsized peptides (∼30 amino acids in length) selected using mRNA display to perform a nucleophilic aromatic substitution reaction (SNAr). Analysis suggested a diverse set of reactive sequences with significant differences in primary sequence, secondary structure and even predicted tertiary structural features. Several of the sequences displayed rapid kinetics allowing for near complete labeling in under one hour. Rosetta ab initio structure prediction of these sequences suggested a landscape of structural features, ranging from beta-sheet-based sequences to those possessing more alpha-helical-like character. Circular dichroism spectroscopy confirmed elements of the structure predictions for the majority of peptides. This analysis additionally uncovered that several peptides underwent secondary structure alterations upon reaction. These results suggest a broad sequence and structural landscape of SNAr active peptides along with a potentially important feature of these biopolymers.



