
In-solution enrichment identifies peptide inhibitors of protein–protein interactions
In-solution enrichment identifies peptide inhibitors of protein–protein interactions
Nature Chemical Biologyvolume 15, pages410–418 (2019)
Fayçal Touti, Zachary P. Gates, Anupam Bandyopadhyay, Guillaume Lautrette & Bradley L. Pentelute
Abstract
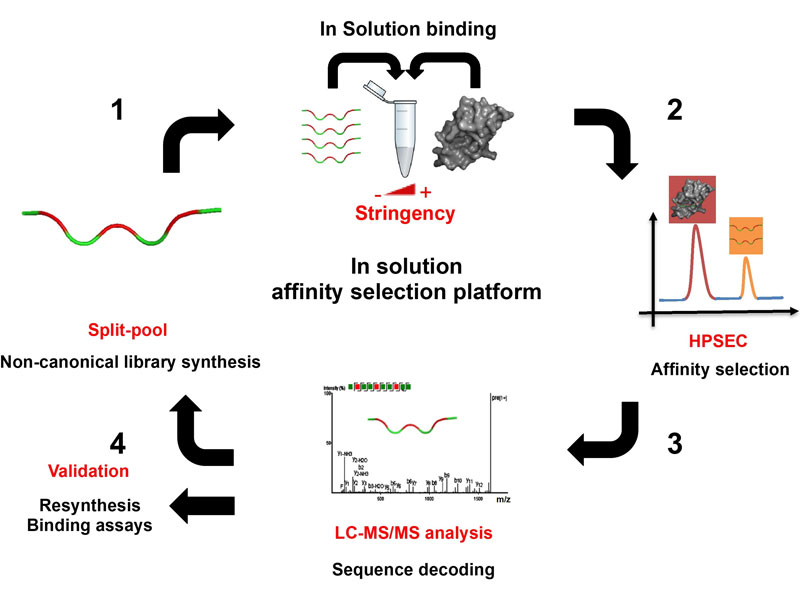
The use of competitive inhibitors to disrupt protein–protein interactions (PPIs) holds great promise for the treatment of disease. However, the discovery of high-affinity inhibitors can be a challenge. Here we report a platform for improving the affinity of peptide-based PPI inhibitors using non-canonical amino acids. The platform utilizes size exclusion-based enrichment from pools of synthetic peptides (1.5–4 kDa) and liquid chromatography-tandem mass spectrometry-based peptide sequencing to identify high-affinity binders to protein targets, without the need for ‘reporter’ or ‘encoding’ tags. Using this approach—which is inherently selective for high-affinity binders—we realized gains in affinity of up to ~100- or ~30-fold for binders to the oncogenic ubiquitin ligase MDM2 or HIV capsid protein C-terminal domain, which inhibit MDM2–p53 interaction or HIV capsid protein C-terminal domain dimerization, respectively. Subsequent macrocyclization of select MDM2 inhibitors rendered them cell permeable and cytotoxic toward cancer cells, demonstrating the utility of the identified compounds as functional PPI inhibitors.



