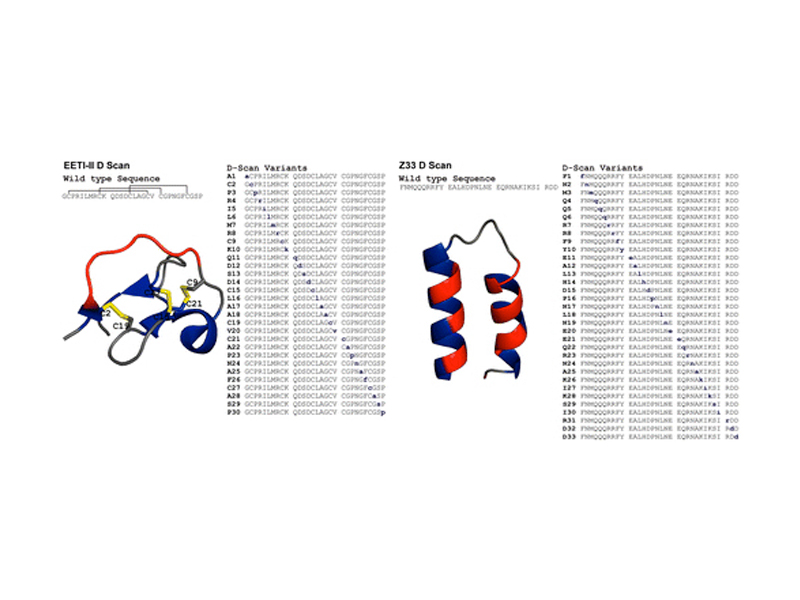
d-Amino Acid Scan of Two Small Proteins
A “D-scan” of two small proteins, the disulfide-rich Ecballium elaterium trypsin inhibitor II (EETI-II) and a minimized Z domain of protein A (Z33), is reported. For each protein, the stereochemistry of one amino acid at a time was inverted to generate a series of diastereomers. In much the same way an alanine scan determines necessary residues for protein function, the D-scan elucidated the critical stereocenters of the 30-residue EETI-II and the 33-residue Z33. The folding properties and activity of each variant were investigated. A total of 24 out of 30 EETI-II D-scan analogues folded to give a three-disulfide product. Of the 24 variants that folded, half were high-affinity trypsin inhibitors, and three were as active as the wild type (WT). Of these 12 active variants, most were substantially less stable to reduction than WT EETI-II (WT first reduction potential −270.0 ± 1.5 mV, WT second reduction potential −307.2 ± 1.1 mV). Similarly, ten Z33 analogues retained high binding affinity to IgG (KD < 250 nM, WT: 24 ± 1 nM) and 12 additional analogues had reduced but appreciable IgG binding affinity (KD between 250 nM and 2.5 μM). As with EETI-II, most Z33 analogues were substantially less stable than the WT (ΔG(H2O, 263 K) = 2.4 ± 1.2 kcal/mol). Collectively, our findings show that the D-scan is powerful new strategy for studying how the stereochemistry of amino acids affects the structure and function of proteins.
Pentelute Lab, MIT, Publications,
15803
wp-singular,portfolio_page-template-default,single,single-portfolio_page,postid-15803,wp-theme-bridge,bridge-core-3.0.1,qode-page-transition-enabled,ajax_fade,page_not_loaded,,paspartu_enabled,paspartu_on_top_fixed,paspartu_on_bottom_fixed,qode_grid_1200,qode_popup_menu_push_text_top,qode-theme-ver-28.7,qode-theme-bridge,disabled_footer_top,wpb-js-composer js-comp-ver-6.8.0,vc_responsive
d-Amino Acid Scan of Two Small Proteins
d-Amino Acid Scan of Two Small Proteins
J. Am. Chem. Soc., 2016, 138 (37), pp 12099–12111
DOI: 10.1021/jacs.6b03765
Publication Date (Web): August 5, 2016
Mark D. Simon†, Yuta Maki†‡, Alexander A. Vinogradov†, Chi Zhang†, Hongtao Yu§, Yu-Shan Lin§, Yasuhiro Kajihara‡, and Bradley L. Pentelute*†
Abstract
A “D-scan” of two small proteins, the disulfide-rich Ecballium elaterium trypsin inhibitor II (EETI-II) and a minimized Z domain of protein A (Z33), is reported. For each protein, the stereochemistry of one amino acid at a time was inverted to generate a series of diastereomers. In much the same way an alanine scan determines necessary residues for protein function, the D-scan elucidated the critical stereocenters of the 30-residue EETI-II and the 33-residue Z33. The folding properties and activity of each variant were investigated. A total of 24 out of 30 EETI-II D-scan analogues folded to give a three-disulfide product. Of the 24 variants that folded, half were high-affinity trypsin inhibitors, and three were as active as the wild type (WT). Of these 12 active variants, most were substantially less stable to reduction than WT EETI-II (WT first reduction potential −270.0 ± 1.5 mV, WT second reduction potential −307.2 ± 1.1 mV). Similarly, ten Z33 analogues retained high binding affinity to IgG (KD < 250 nM, WT: 24 ± 1 nM) and 12 additional analogues had reduced but appreciable IgG binding affinity (KD between 250 nM and 2.5 μM). As with EETI-II, most Z33 analogues were substantially less stable than the WT (ΔG(H2O, 263 K) = 2.4 ± 1.2 kcal/mol). Collectively, our findings show that the D-scan is powerful new strategy for studying how the stereochemistry of amino acids affects the structure and function of proteins.
Category
2016, Publications